
I’ve been thinking about some new iterations of The Synthetic Kingdom, from all the feedback I’ve had on the project since last year.
I presented this work at the ‘Beyond the Tree of Life Meeting’, in London, 10 July 2009.
How will we classify what is natural or unnatural when life is built from scratch?
Synthetic Biology is turning to the living kingdoms for its materials library. No more petrochemicals: instead, pick a feature from an existing organism, locate its DNA code and insert it into a biological chassis. From DIY hacked bacteria to entirely artificial, corporate life-forms, engineered life will compute, produce energy, clean up pollution, make self-healing materials, kill pathogens and even do the housework.
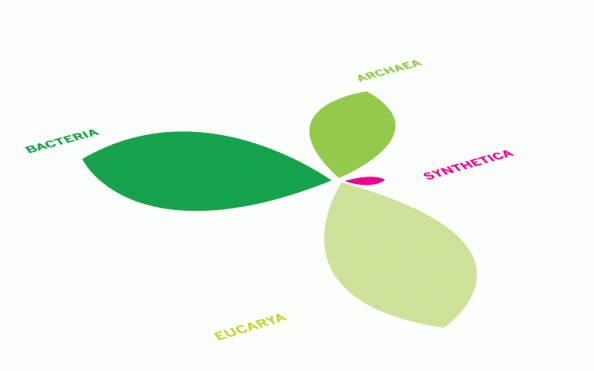
The Tree of Life is always changing, ever since we first created it. Now, we are adding to the living kingdoms for the first time. But these synthetic organisms are no different to other life forms, except that we invented them.
We will have to insert an extra branch into the Tree of Life. The Synthetic Kingdom is part of our new nature.




Center for Genomic Gastronomy
July 15, 2010
Looks good.
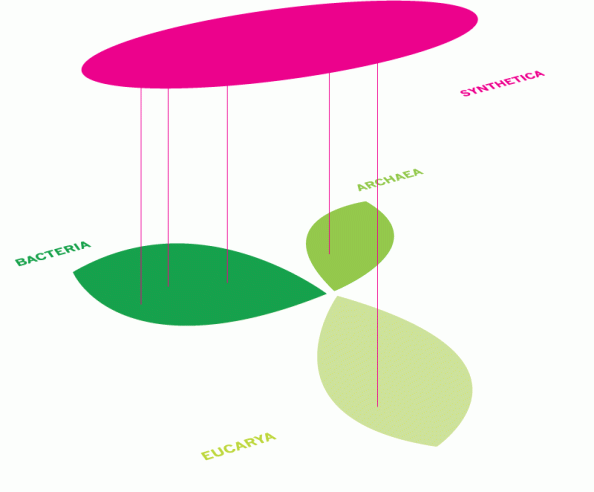
The Center for Genomic Gastronomy has taken what looks to be some combination of the “weeds” and “network spaghetti” model to map out the provenance of the transgenic ingredients we are studying.
For Genetically Engineered organisms (as opposed to synth bio, etc.) it may be useful to put a weed close to the “main” organism and then a network connector to the gene that was inserted. I am wondering if each phylum under the synthetic kingdom needs its own visualization technique in order to appropriately visualize the process by which it was generated?
Good luck on further iterations!
alexandradaisy
July 15, 2010
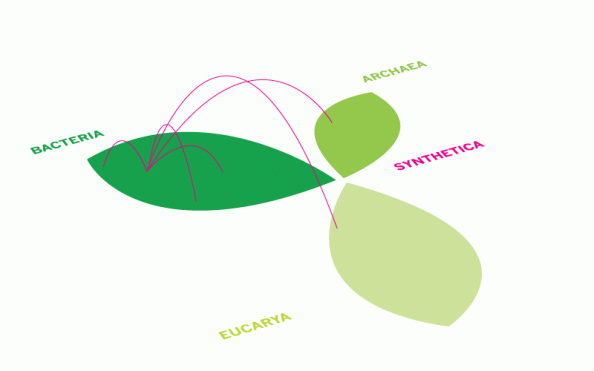
Thanks for the comment! The overall consensus at the Tree of Life meeting was in agreement, heading towards a Spaghetti Network for the Synthetic Kingdom. That said, there was much debate about the Tree of Life itself, which looks like its status is fragile. I’m suspecting soon we’ll have a Fuzzball of Life or a Pompom of Life (in 3D, but is rooted, unlike the fuzz network), with a Synthetic Spaghetti Network coming off it. Next on the drawing list…
I’ll post some of the suggestions soon. In the meantime, here are some of the speakers’ suggestions: http://www.flickr.com/photos/alexandradaisy/4780383488/
Lab Rat
July 22, 2010
Nice work! I like the synthetic kingdom as little branching weeds, that works best I think until someone (probably Venter…) actually makes an empty synthetic chases to put genes into 🙂
Center for Genomic Gastronomy
August 9, 2010
One of the values of the “Tree” metaphor is about mapping, geography, distance and direction. Was there any discussion about the 2-D Base map that would underly the 3-D map? A strong base map could help the spaghetti contain directional / distance information. Also, what about organisms that have more than one transgene? Would a bigger prototypical change for one transgene as opposed to another have a thicker line? Great investigations. If you have ever edited wikipedia, I would love to see this debate controversy be archived there in some manner. Also, do you know of any trees of life that include agricultural cultivars, or is that just too obscure? Great stuff!
Rich Pell
August 9, 2010
Hey Daisy, these look great!
I agree about the “spaghetti map”. When we started mapping the transgenic tree of life, it seemed really important that the mapping occur at the transgene level so that, like Zack said, you can show the relationship of more than one transgene at a time. The problem is that this works great for mapping out a single GMO’s relationship to the tree of life, but becomes very cluttered when mapping out these relationship at a full tree of life level. Still, interesting and recognizable patterns emerge: The model organisms show up as major hubs of GMO’s as well as sources of transgenes for other GMO’s. We ended up using two different tree depictions, one for individual GMO’s and one for the global tree. The individual ones include the tree structure, with the evolutionary relationships between the host and the donor genes present. The global one did not include the evolutionary relationships and gave primacy instead to the transgene connections.
It’s fun problem. I wish I could have joined you at “Beyond the Tree of Life”!
Paul Record
September 11, 2015
Alexandra some very interesting ideas. I love the ideas around synthetics biology it could evolve very differently from existing biology. Could artificisl intellegence evolve from it. As for classification development you like to have a look at protege from Stanford uni as a way to develop an ontology for synthetica. It uses a symantec web language OWL. Regards Paul